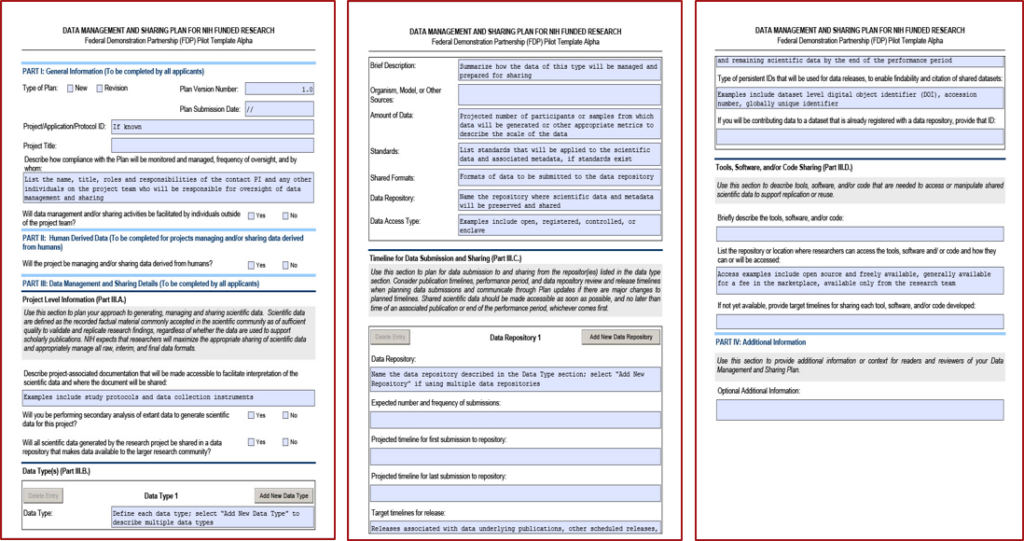
The National Institutes of Health (NIH) requires a Data Management and Sharing Plan (DMSP) in all competing applications that generate scientific data. A DMSP should cover six key elements as outlined in NOT-OD-21-014. While not required, using a DMSP template helps researchers address all six elements and sub-elements required by the NIH DMS policy. The most widely used DMSP template is the optional DMSP format page (Fig 1). However, two updated pilot templates, Alpha (Fig 2) and Bravo (Fig 3) were released in 2023 with the opportunity to provide feedback to the NIH (NIH Extramural Nexus announcement on November 30, 2023). This blog will provide practical tips for writing a strong DMSP based on more than 100 submitted DMSPs and public feedback from four NIH Institutes (NICHD, NCI, NIMH, and NIBIB). I’ll provide general tips as well as element-specific guidance to help you write successful DMSPs for your NIH applications.


General Tips
Tip #1: First, determine whether the NIH Genomic Data Sharing (GDS) policy applies to your study, including IC-specific thresholds (e.g. NCI thresholds). Second, determine if your research is considered human subjects research using decision tools (Decision Tool, Flowchart, Infographic). Finally, determine if your research involves secondary analysis. Answering these questions helps NIH staff review your DMSP and sets a framework for your DMSP because it affects which data/when/where/how to share. If you use the 2023 FDP Pilot DMSP templates, these questions are asked upfront, but if using the 2022 optional format page, add this information in “Element 1”.
Tip #2: Create a table listing all data types in your research strategy plan. It is important to list “all” data types, not just selective data types you think are sharable. For each data type, include a brief description that includes: the source (human, animal, cell lines, existing datasets for secondary analysis, etc.), shared data file format, amount (file size in GB/TB or the number of animals or human participants), and the designated repository for your data type. Although NOT-OD-21-014 doesn’t explicitly require a data source, recent feedback from NIH ICs suggests including a source since it affects subsequent DMSP elements. The data table should be added in Element 1A of the Optional Format page or enter the information into the existing table in the Alpha or Bravo templates.
Tip #3: Identify the appropriate data repositories that meet the desirable characteristics of data repositories for federally funded research for each data type. Discuss repository options with your program officer (PO) if uncertain. Utilize the decision tree for repository selection and explore resources for finding a repository. WashU offers two institutional data repositories that can accept all data types: Digital Commons Data@Becker, managed by Becker Medical Library for the School of Medicine, and the WashU Research Data Repository (WURD), managed by University Libraries for those on the Danforth Campus.
NIH Optional DMSP Format Page Guidance
Element 1: Data Type
A) Types and amount of scientific data expected to be generated in the project:
As mentioned in Tip #1, determine whether the NIH Genomic Data Sharing (GDS) policy applies to your study, then state whether your project involves human subjects research and secondary analysis. Then organize your data types using a table, as described in Tip #2. The NIH also expects you to describe where each data type will be managed before it is placed in a public repository for sharing. This is especially important if you use a local storage platform that incurs a cost (e.g., RIS, Data Lake, Center for Genome Sciences Computational Cluster, WUBIOS facility, etc.). Local data management considerations are an allowable DMS cost, but the DMS budget justification must be based on your DMSP, so please ensure that your DMS budget justification aligns with the information provided in your DMSP!
B) Scientific data that will be preserved and shared, and the rationale for doing so:
Clearly state which data will be preserved and shared and justify any limitations on data sharing. SBIR/STTR applicants are permitted to withhold applicable data for up to 20 years after the award date, by including this NIH recommended statement. If you are doing secondary analyses on previously shared data, you do not need to reshare the existing, shared primary data, but any new, derived data generated as a result of secondary analyses are expected to be shared, unless the use of data obtained from repositories or other sources and derived data is subject to limitations on sharing as a condition of access. Not all data generated during NIH-supported research will constitute scientific data under the DMS Policy. Please see the example exclusion criteria on the FAQ page. Also, the NIH acknowledges that data sharing may have limitations and provides examples of justifiable reasons for restricting sharing. Please cite specific reasons (e.g., GDPR requirements or specific laws) rather than using vague language such as “legal” considerations.
C) Metadata, other relevant data, and associated documentation:
Describe any additional information necessary for others to interpret your data. Examples of associated documentation include study protocols, data dictionaries, survey instruments, and readme files. Data-type-specific metadata are typically collected by domain-specific repositories (e.g., ImmPort Data Model, GEO etc.) via sample submission templates, so be prepared to provide this metadata for each data type. If you are using generalist repositories, including WashU institutional data repositories (Digital Commons Data@Becker or WURD), they typically use the DataCite metadata schema to collect searchable study-level metadata. If you select Digital Commons Data@Becker for your data types, please refer to the DMSP Template for Digital Commons Data@Becker.
Element 2: Related Tools, Software, and/or Code
This section focuses on what will be required for “others” to open your data files and perform further analysis. This may not be the same tools “you” used to collect and process the data. It is recommended that you convert your data files from proprietary formats to open file formats before placing them in a data repository (See Element 3 below) so that others can use publicly available open-source software to access and further process your data.
If custom code is necessary to reproduce your results, make it publicly available in a repository like GitHub. Mention the availability of your code in this section and note that the GitHub repository will be archived with Zenodo to obtain a DOI for citation, which can be included in your publications, RPPRs, and Biosketch.
Element 3: Standards
This element focuses on achieving the interoperability of FAIR data principles. To ensure interoperability, convert your shared files into open file formats and adhere to community-accepted standards and ontologies. Before concluding that there are no widely accepted standards for your data types, please check for standards on fairsharing.org, the largest database for standards and ontologies. Also, the NIDDK Data and Metadata Standards Examples for DMS Plans and NICHD Data Standards Resources can guide you in writing this section.
Element 4: Data Preservation, Access, and Associated Timelines
A) Repository where scientific data and metadata will be archived:
If you have included repositories for each data type in the table in Element 1A, you can simply state that data will be archived in the repositories mentioned in Table 1. If not, you can list the names of domain-specific data repositories for each data type or a generalist repository that can accept all data types.
B) How scientific data will be findable and identifiable:
Your data will be findable and identifiable via the Persistent Unique Identifiers (PIDs) assigned to your dataset by the data repositories (e.g., DOIs, Accession numbers).
C) When and how long the scientific data will be made available:
The duration that your data will be accessible depends on the retention policy of the data repositories you choose. Ensure that you choose a data repository that meets the minimum data retention period required by the NIH (3 years) and your institution (6 years for WashU). In addition, the NIH DMS policy requires that data should be shared at the time of associated publications or the end of the award period, whichever comes first. However, if your study is subject to the NIH GDS Policy, consult the expectations outlined in the “Data Submission and Release Expectations for Genomic Data” guidance and include them in this section.
Element 5: Access, Distribution, or Reuse Considerations
A) Factors affecting subsequent access, distribution, or reuse of scientific data:
If your data is subject to patent applications or license agreements or if your project involves human subjects research, several potential factors might affect the subsequent access, distribution, or reuse of your data. Check potential justifiable factors listed by the NIH and confirm those that apply to your data with the Joint Research Office for Contracts (JROC) and the Human Research Protection Office (HRPO). If you are generating whole genome sequencing from HeLa cells, check the NIH sample plan on HeLa Cell WGS and follow the guidelines in NOT-OD-24-098. If your study is subject to the NIH GDS policy, indicate if your study should be designated as “sensitive” for the purposes of access to Genomic Summary Results (GSR), as described in NOT-OD-19-023.
B) Whether access to scientific data will be controlled:
If your data is generated from non-human sources, the NIH expects data to be shared openly without access restrictions. If your data is human-derived, check with HRPO/IRB to confirm which data can be shared (de-identified individual participant data vs. aggregated summary data) and the appropriate level of access (open vs. controlled).
C) Protections for privacy, rights, and confidentiality of human research participants:
Outline the measures you will implement to protect the privacy, rights, and confidentiality of human participants. These steps might involve de-identifying data by removing personally identifiable information (PII), obtaining a certificate of confidentiality, or implementing other suitable protective measures like access control and HIPAA training. When addressing this section, adhere to the guidelines provided in NOT-OD-213 and NOT-OD-214.
Element 6: Oversight of Data Management and Sharing
In this section, various NIH Sample DMSPs discuss institutional oversight. However, it is important to clarify that NIH does not currently mandate institutional oversight. Instead, sponsoring institutions have the flexibility to handle oversight based on their individual circumstances.
At WashU, the Office of Sponsored Research Services (OSRS) has addressed this matter by assigning data management responsibility and sharing oversight to Principal Investigators (PIs). PIs are responsible for ensuring compliance with the approved DMSP.
For WashU investigators seeking guidance, sample language for Element 6 can be found in the Example DMS Plans in the ICTS WUSTL Grants Library or DMSP Template for Digital Commons Data@Becker.
Finally, before you write a DMSP from scratch, be sure to check out Sample DMS Plans on the BeckerDMS website, which include NIH sample DMS Plans, examples DMS Plans in the WUSTL Grants Library (Submitted or Awarded), and Featured DMS Plans in DMPTool. These samples provide a great starting point to create a custom DMSP for your project.
I hope you find these tips helpful! If you have any questions or would like personal assistance in reviewing your DMSP, please email seonyoung.kim@wustl.edu or request a consultation via the BeckerDMS website.
Resources
- NIH Data Management and Sharing Policy
- DMSP Template for Digital Commons Data@Becker
- NOT-OD-21-014 (Elements of an NIH Data Management and Sharing Plan)
- FDP Data Management and Sharing Town Hall #3 (Video, Slide Deck, Transcript): Feedback from four NIH ICs (NICHD, NCI, NIMH, and NIBIB)
- NIDDK Data and Metadata Standards Examples for DMS Plans
- NICHD Data Standards Resources
- ImmPort Data Model
- NOT-OD-24-098 (Update to Hela Cell Whole Genome Sequence Data Submission and Access Under the NIH Lacks-Family Agreement)
- NOT-OD-22-214 (Supplemental Information to the NIH Policy for Data Management and Sharing: Responsible Management and Sharing of American Indian/Alaska Native Participant Data)
- NOT-OD-22-213 (Supplemental Information to the NIH Policy for Data Management and Sharing: Protecting Privacy When Sharing Human Research Participant Data)
